The Photophysics of Green Fluorescent Protein: Influence of the Key Amino Acids at Positions 65, 203, and 222: Biophysical Journal

Precise determination of protein extinction coefficients under native and denaturing conditions using SV-AUC | SpringerLink

Prediction of Molar Extinction Coefficients of Proteins and Peptides Using UV Absorption of the Constituent Amino Acids at 214 nm To Enable Quantitative Reverse Phase High-Performance Liquid Chromatography−Mass Spectrometry Analysis | Journal
![PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/af80a5d2beee749cff2f5d98dc4a8f0dfdd48074/3-Table2-1.png)
PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar
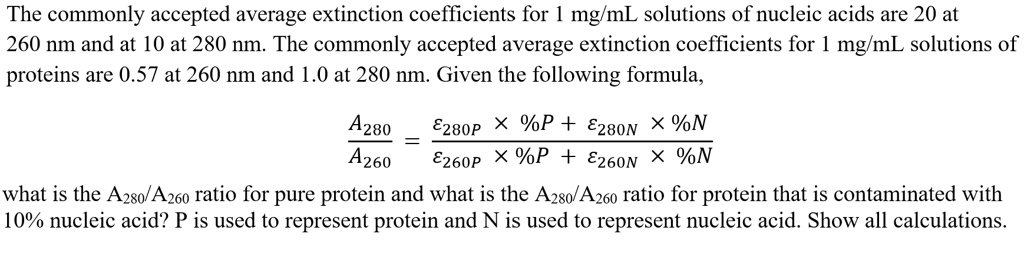

SOLVED: The commonly accepted average extinction coefficients for mg/mL solutions of nucleic acids are 20 at 260 nm and at 10 at 280 nm The commonly accepted average extinction coefficients for mglmL
Engrailed and Hox homeodomain proteins contain a related Pbx interaction motif that recognizes a common structure present in Pbx. - Abstract - Europe PMC

Impact of external amino acids on fluorescent protein chromophore biosynthesis revealed by molecular dynamics and mutagenesis studies - ScienceDirect
![PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/af80a5d2beee749cff2f5d98dc4a8f0dfdd48074/2-Table1-1.png)
PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar

SOLVED: It is possible to estimate the molar extinction coefficient of a protein from knowledge of its amino acid composition, as shown from your experiences with ExPASY: From the molar extinction coefficient